Code
library(tidyverse)
library(here)
library(janitor)
library(NbClust) #cluster package
library(cluster)
library(dendextend)
library(ggdendro)
library(factoextra)February 14, 2024

Data Description:
The data used in this analysis is a data package from Santa Barbara Coastal (SBC) Long Term Ecological Research (LTER). It contains stream water chemistry measurements taken in various watersheds throughout the Santa Barbara area, with data collection beginning in 2000. As described by the data source, this “dataset is ongoing, and data will be added approximately annually. Stream water samples are collected weekly during non-storm flows in winter, and bi-weekly during summer. During winter storms, samples are collected hourly (rising limb) or at 2-4 hour intervals (falling limb). Analytes sampled in the SBC LTER watersheds include dissolved nitrogen (nitrate, ammonium, total dissolved nitrogen); soluble reactive phosphorus (SRP); particulate organic carbon, nitrogen and phosphorus; total suspended sediments; and conductivity.”
Data Citation:
Santa Barbara Coastal LTER and J. Melack. 2019. SBC LTER: Land: Stream chemistry in the Santa Barbara Coastal drainage area, ongoing since 2000 ver 16. Environmental Data Initiative. https://doi.org/10.6073/pasta/67a558a24ceed9a0a5bf5e46ab841174
Objective:
This analysis aims to use complete linkage agglomerative hierarchical clustering to compare water chemistry by specific sites in the SBC LTER dataset. This “bottom-up” approach to clustering will allow us to group sites whose water quality are more similar to each other and different from other sites in the dataset.
Pseudo-code:
summary() function to identify all columns that have greater than 50% NA values and remove them.na.rm = TRUE when summarizing to remove any rows with NA’s.stats::hclust() function to perform the complete linkage agglomerative hierarchical clustering.Process to Deal w/ NAs:
summary() function, and drop them from the analysis.na.rm = TRUE when summarizing water quality measurements at each site to remove all other NA values remaining in the data. I chose this method instead of using drop_na() for the entire data frame because this removed NAs for each respective measurement when calculating their mean values rather than the entire row of data wherever an NA value was present.# examine which columns have more than 50% NA values
summary(stream_chem_raw) ## 'tpc_uM' ; 'tpn_uM' ; 'tpp_uM ; 'tss_mgperLiter'
# drop columns with > 50% NA's from the df
stream_chem_df <- stream_chem_raw %>%
select(site_code, nh4_uM, no3_uM, po4_uM, tdn_uM, tdp_uM, spec_cond_uSpercm)
# make a data frame with a single summary row per site
# take the mean of all observations at each site
# drop NAs using `na.rm = TRUE` when summarizing
### use this df when calculating the Euclidian distance ###
stream_chem_means <- stream_chem_df %>%
group_by(site_code) %>%
summarize(nh4_avg = mean(nh4_uM, na.rm = TRUE),
no3_avg = mean(no3_uM, na.rm = TRUE),
po4_avg = mean(po4_uM, na.rm = TRUE),
tdn_avg = mean(tdn_uM, na.rm = TRUE),
tdp_avg = mean(tdp_uM, na.rm = TRUE),
spec_avg = mean(spec_cond_uSpercm, na.rm = TRUE))
# scale the average measurements at each site
stream_mean_scale <- stream_chem_means %>%
select(-site_code) %>%
scale()
### If I want to drop all NA's instread of using na.rm in summarize() ###
# # drop all rows with NA's
# stream_complete <- stream_chem_df %>%
# drop_na()
#
# # scale the measurements at each site
# stream_scale <- stream_complete %>%
# scale() # this function centers and/or scales the columns of a numeric matrix
#
# # compare the scaled and complete df's
# summary(stream_complete)
# summary(stream_scale) # the mean measurement at each site is 0 ### complete ###
# calculate Euclidean distance; should this be using the mean stream df???
stream_dist <- dist(stream_mean_scale, method = "euclidean")
# Hierarchical clustering (complete linkage)
stream_hc_complete <- hclust(stream_dist, method = "complete")
# # Plot it (base plot):
# plot(stream_hc_complete, cex = 0.6, hang = -1)# prep vectors for labelling
stream_sites <- c("AB00", "AT07", "BC02", "DV01", "GV01", "HO00", "MC00",
"MC06", "ON02", "RG01", "RS02", "SP02", "TO02")
int_placeholders <- c(4, 3, 2, 1, 7, 12, 5, 8, 13, 10, 9, 6, 11)
# make the dendrogram
ggdendrogram(stream_hc_complete) +
theme_classic() +
scale_x_continuous(labels = stream_sites, breaks = int_placeholders) +
labs(x = "Measurement site",
y = "Distance between clusters")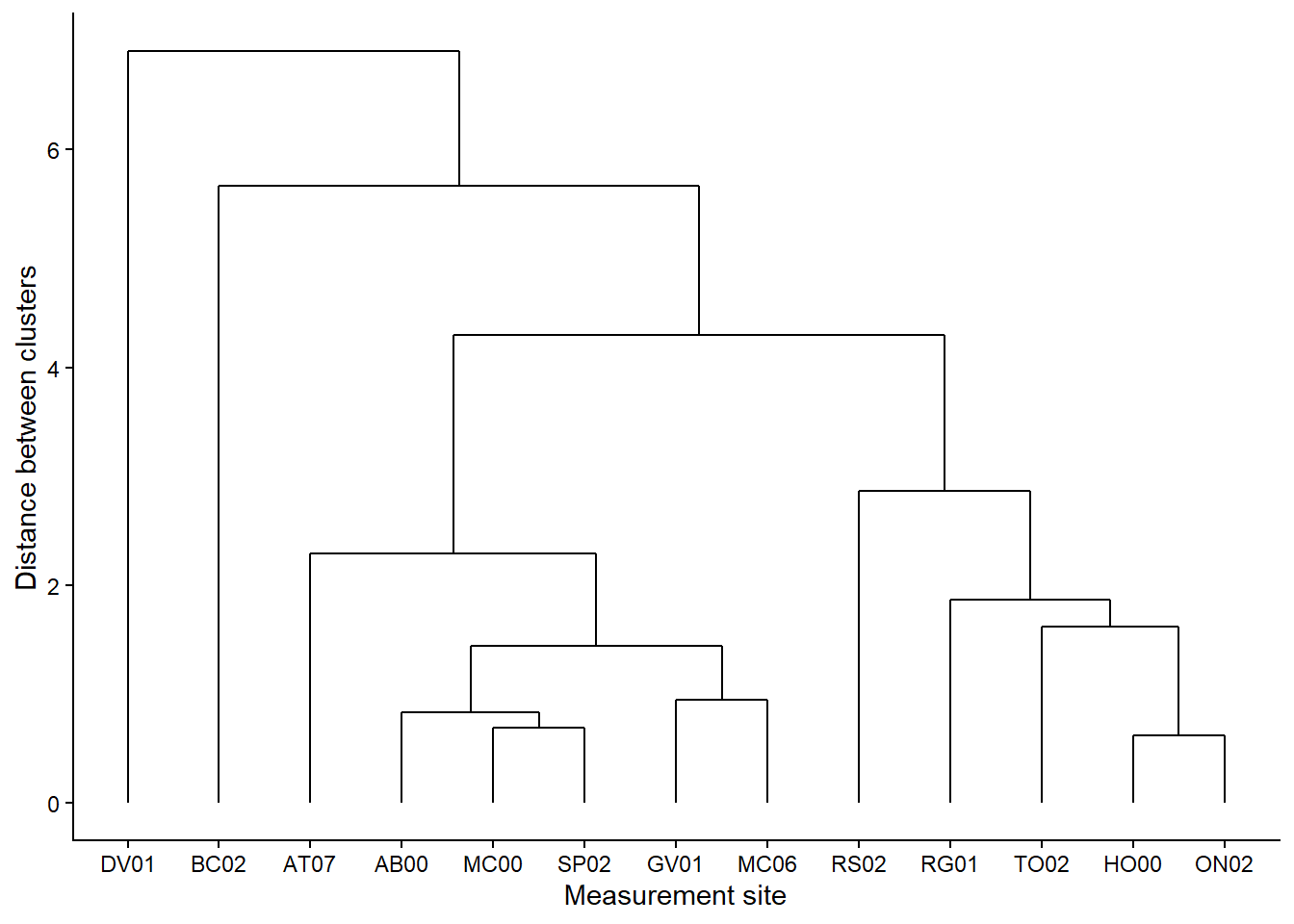
This assignment was created and organized by Casey O’Hara at the Bren School for ESM 244 (Advanced Data Analysis for Environmental Science & Management). ESM 244 is offered in the Master of Environmental Science & Management (MESM) program.
@online{pepperdine2024,
author = {Pepperdine, Maxwell},
title = {Comparing Water Chemistry in {Santa} {Barbara} Streams with a
Cluster Analysis},
date = {2024-02-14},
url = {https://maxpepperdine.github.io/posts/2024-02-14-hierarchical-cluster/},
langid = {en}
}